Public-health and biomedical decisions increasingly depend on converting genomic data into actionable evidence. I develop and implement genomic methods that bridge research and real-world use, spanning culture-free TB sequencing and resistance prediction, respiratory pathogen surveillance, and vector and malaria population genomics.
My work focuses on turning pathogen and vector genomics into decision-grade inputs for surveillance, intervention planning, and disease control—across both pharma / real-world evidence and global health implementation contexts.
Applied across:
- Pharma & RWE: resistance biomarkers, genomic surveillance, post-licensure monitoring
- Global health: national surveillance systems, vector control strategy, TB and malaria program support
What I Build
My work centres on operational systems, not isolated analyses:
- End-to-end genomic surveillance pipelines (sample → sequence → insight)
- Culture-free and rapid resistance diagnostics
- Population-genetic inference for intervention planning
- Scalable analytics for real-world and public-health data
Where This Fits in Decision-Making
Across projects, genomics is used to support decisions such as:
- Faster treatment selection through resistance prediction
- Improved timing and targeting of vaccines, mAbs, and vector control
- Evaluation of program impact and surveillance effectiveness
- Translation of genomic data into policy-relevant evidence
Portfolio: Selected Works:
- Culture-free TB whole-genome sequencing and drug resistance profiling
- RSV genomic surveillance in a hospital-linked system
- Malaria transmission network inference
- Global migration and insecticide resistance in Aedes aegypti
Contact
I’m interested in roles and collaborations spanning pharma / biotech, real-world evidence, and global health, particularly where genomics informs real-world decisions.
seth.redmond@yale.edu / seth.redmond@gmail.com
Publications & Links
- GitHub: [https://github.com/sethnr]
- Google Scholar: [https://scholar.google.com/citations?user=Kty5i3wAAAAJ]
- LinkedIn: [https://www.linkedin.com/in/sethnr/]
Culture-free TB whole-genome sequencing and drug resistance profiling
Context
Tuberculosis sequencing remains slow and laborious. Reliable sequencing typically requires intermediate culturing, which introduces significant delay and is poorly suited to in high-burden settings, limiting the use of genomics for timely treatment decisions.
What I built
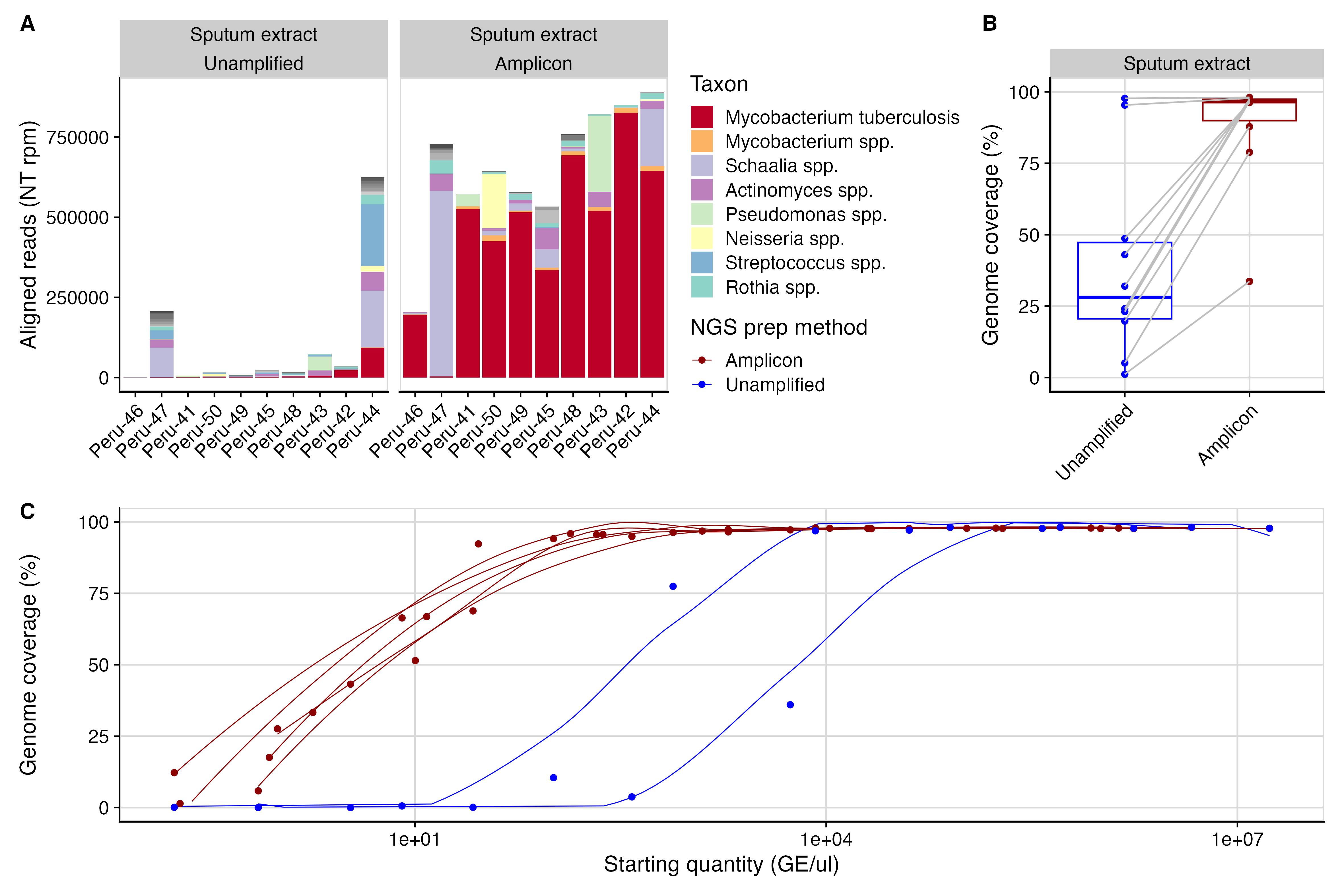
A tiled amplicon sequencing approach for Mycobacterium tuberculosis that enables whole-genome sequencing directly from patient sputum, without an intermediate culture step, enabling rapid comprehensive resistance profiling.
Why it matters
- Generates high genome coverage from uncultured sputum, from even highly metagenomic samples
- Shortens time from sample to drug resistance profile, and could enable treatment tailoring
- Supports scalable, programmatic drug-resistance surveillance
Links
RSV genomic surveillance in a hospital-linked system
Context
With the rollout of RSV monoclonal antibodies and vaccines, the limiting factor is no longer the availability of interventions, but the ability to target them effectively. Local transmission dynamics and age-structured spread are poorly resolved by routine surveillance.
What I built
An end-to-end RSV genomic surveillance system integrated with a major hospital network, linking viral sequencing with epidemiological metadata from routine clinical sampling.
Why it matters
- Resolves fine-scale, local RSV transmission patterns
- Provides age-specific insights with direct relevance to mAb and vaccine targeting
- Demonstrates a scalable model for linking clinical care and genomic surveillance
Links
Global migration and insecticide resistance in Aedes aegypti
Context
Vector control strategies often assume geographically stable and genetically homogeneous mosquito populations, despite differences in insecticide resistance, host preference, and vector capacity. Aedes is known to exhibit two subspecies, with the ancestral African form thought to have a lower capacity to transmit arboviruses than the derived global form.
What I built
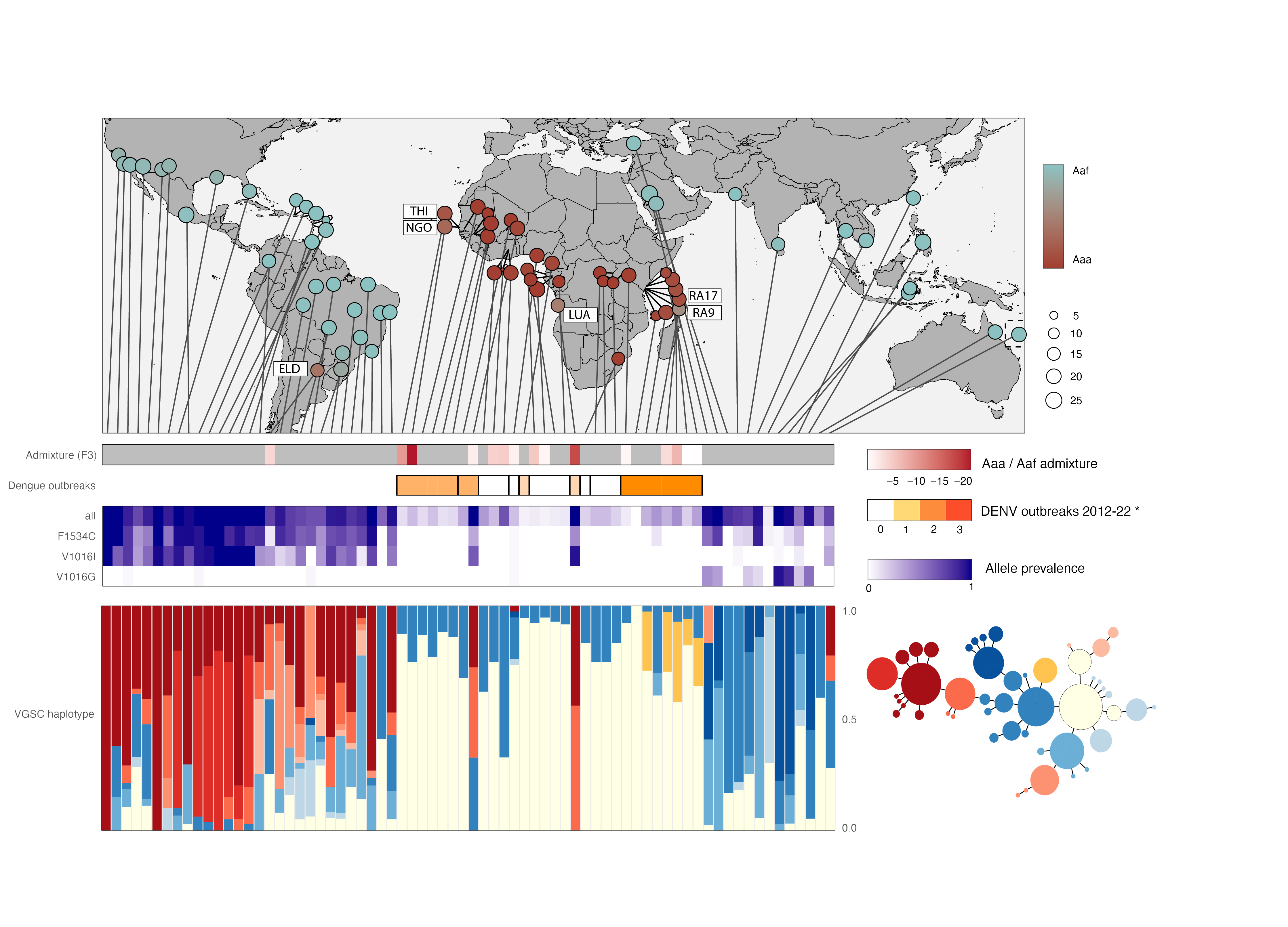
Population admixture analyses between African and global populations of Aedes aegypti to resolve contemporary migration patterns, track the global spread of pyrethroid resistance alleles, and identify regions of reinvasion that could affect dengue transmission.
Why it matters
- Reveals extensive contemporary mosquito migration, explaining the rapid dissemination of resistance variants
- Admixed mosquito populations are found in African regions with the heaviest dengue burden.
- Informs resistance management, insecticide policy, and intervention planning
Links
Malaria transmission network inference
Context
Almost all genomic epidemiology methods for P. falciparum rely on pairwise relatedness, however as malaria transmission declines, these methods become less applicable, just at the point where elimination efforts depend on identifying who is infecting whom.
What I built
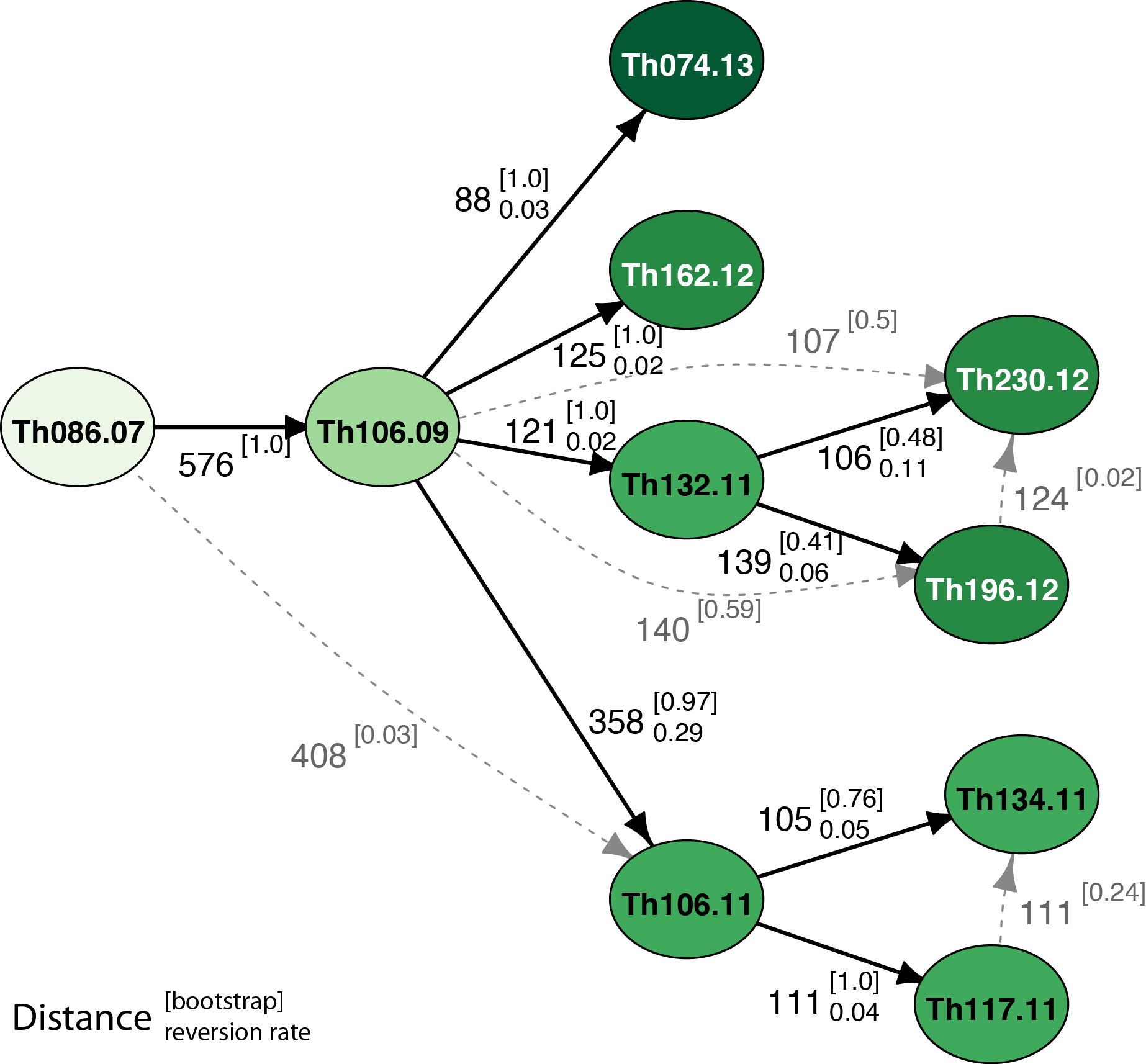
Genomic epidemiology methods combining wet-lab and computational approaches to increase variant calling fidelity and using genetic distance from de novo mutations to infer individual-level transmission networks for Plasmodium falciparum.
Why it matters
- Enables reconstruction of transmission networks at fine scale
- Shows the independent transmission of multiple clonal lineages through the dry season
- Illustrates for the first time that genetic distance can be used for epidemiological inference in Plasmodium falciparum
Links
github / Redmond et al, Mol Bio Evo 2018
What I Build
My work centres on operational systems, not isolated analyses:
- End-to-end genomic surveillance pipelines (sample → sequence → insight)
- Culture-free and rapid resistance diagnostics
- Population-genetic inference for intervention planning
- Scalable analytics for real-world and public-health data
Where This Fits in Decision-Making
Across projects, genomics is used to support decisions such as:
- Faster treatment selection through resistance prediction
- Improved timing and targeting of vaccines, mAbs, and vector control
- Evaluation of program impact and surveillance effectiveness
- Translation of genomic data into policy-relevant evidence
Publications & Links
- GitHub: [https://github.com/sethnr]
- Google Scholar: [https://scholar.google.com/citations?user=Kty5i3wAAAAJ]
- LinkedIn: [https://www.linkedin.com/in/sethnr/]